―Combining molecular simulations and protein language models to augment training data―
Researchers) DEGUCHI Teppei, Research Assistant, KOBAYASHI Kaito, Researcher, SAITO Yutaka, Joint Appointed Fellow, Artificial Intelligence Research Center
- For machine learning based protein functional value predictions, functional values are computed using molecular simulations and protein language models to be utilized as pseudo-training data.
- Achieving highly accurate prediction of protein functional values even with limited experimental data
- Enables efficient development of functional proteins
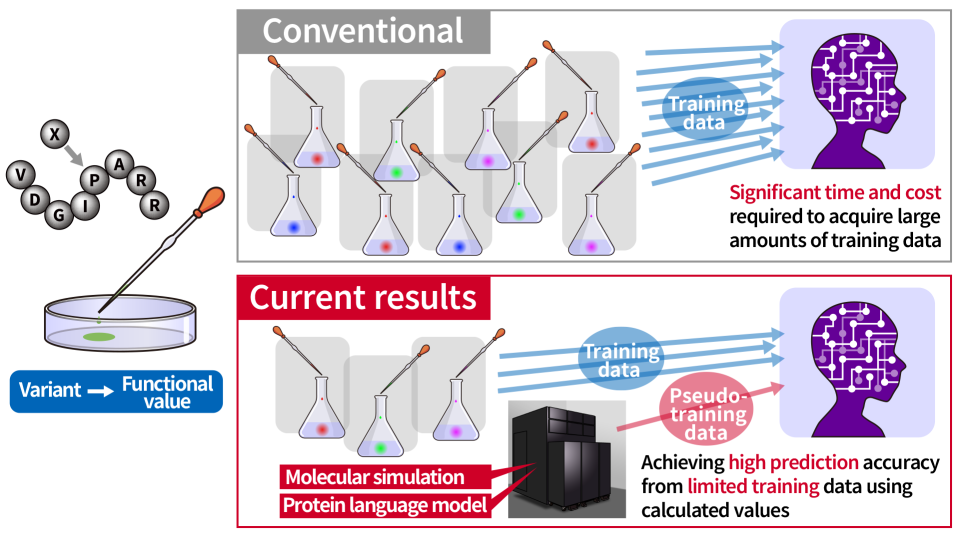
Improving the accuracy of protein functional value prediction through data augmentation
Proteins are biomolecules that play various functional roles within living organisms. For example, they can catalyze chemical reactions as enzymes or recognize specific molecules as antibodies. These functions are also widely utilized in industrial and medical fields. Proteins are composed of amino acids that are connected to each other in a chain-like structure. Modifying this sequence can create functional proteins with improved properties such as activity and stability.
In recent years, progress has been made in developing machine learning based methods for predicting protein function to efficiently design functional proteins with desired properties. Specifically, models are trained using experimental observations of functional values as training data to optimize models to predict functional values for given amino acid sequences. However, making accurate predictions requires large-scale experimental data for training purposes, which is costly in terms of both time and materials.
One effective means of overcoming this problem is data augmentation. This approach involves supplementing model training with computational values obtained through simulations and so on, which are then used as pseudo-training data alongside experimental data. While this approach has previously been applied to predicting protein stability, designing proteins with desired functions requires extending it to properties such as protein-protein binding affinity, enzyme activity, cytotoxicity, and fluorescence intensity, in addition to stability.
Researchers at AIST have developed a machine learning approach that uses molecular simulations and protein language models to accurately predict protein functional values from a small amount of experimental data.
In recent years, researchers have increasingly used machine learning methods to predict protein functions for designing functional proteins. However, this requires significant time and material costs due to the need for large amounts of experimental observations as training data. As a result, methods that use computational values for pseudo-training data along with experimental data have gained attention. Although these methods were previously used to predict protein stability, expending their scope to include predictions of binding affinity and enzyme activity is necessary to design functional proteins tailored to specific purposes.
We have developed a novel method to predict protein functional values. This method takes advantage of functional values computed via molecular simulation and protein language models as pseudo-training data. It achieves high accuracy in predicting functional value even with limited experimental data. Furthermore, we have expanded its applicability beyond protein stability to include binding affinity, enzyme activity, cytotoxicity, and fluorescence intensity. This achievement enables more efficient development of functional proteins compared to existing methods.
Journal:Briefings in Bioinformatics
Title of paper:Data-efficient protein mutational effect prediction with weak supervision by molecular simulation and protein language models
Authors:Teppei Deguchi, Nur Syatila Ab Ghani, Yoichi Kurumida, Shinji Iida, Kaito Kobayashi, Yutaka Saito
DOI:10.1093/bib/bbaf536