Researchers) Molecular Biosystems Research Institute, MORIYAMA Minoru, Senior Researcher, KOGA Ryuichi, Chief Senior Researcher, FUKATSU Takema, Principal Researcher
- Loss-of-function of tryptophan-degrading enzyme transforms E. coli and related bacteria into stinkbug symbionts
- The enzyme gene is absent in natural stinkbug symbionts suggesting that the gene loss is involved in symbiotic evolution
- A key molecular basis for symbiotic evolution is elucidated by integrating laboratory evolution experiments with field population surveys
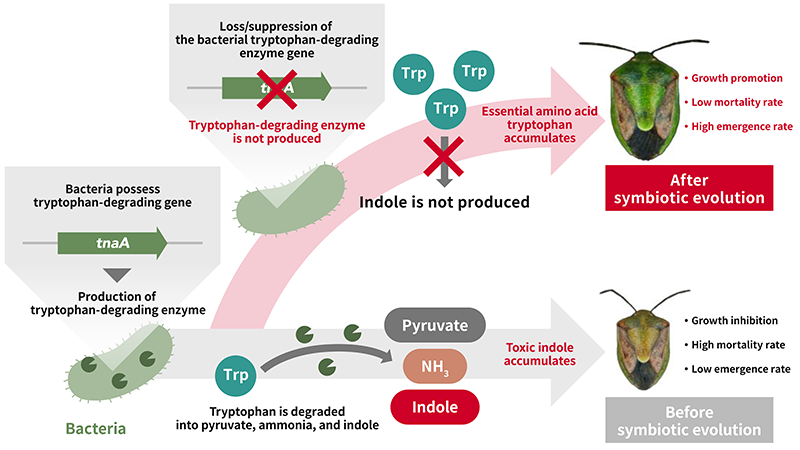
Molecular mechanism as to how E. coli evolves into stinkbug symbiont
In the ecosystem, diverse organisms do not live in isolation. Instead, they often form symbiotic relationships with other organisms to adapt to their respective habitats. For example, insects, which constitute the majority of biodiversity in the terrestrial ecosystem, are often in close symbiotic relationship with microorganisms. These symbiotic microbes are known to influence their hosts’ survival, growth, and biological functions. In the medical field, the human gut microbiota has been reported to influence not only digestive physiology and disease but also various health conditions ranging from constitution to mental health. This has attracted significant attention. In this context, gut symbiotic bacteria have drawn considerable interest in recent years from the perspective of elucidating and utilizing novel biological functions derived from the biodiversity.
AIST researchers discovered that the loss of function of a single enzyme gene encoding tryptophanase transforms E. coli into a gut bacterial symbiont that supports the growth and survival of the host insect. This discovery was made using the uniquely developed experimental symbiotic evolution system between the brown-winged green stinkbug Plautia stali and E. coli. The loss of the tryptophan-degrading enzyme resulted in an increased concentration of the essential amino acid tryptophan and a decreased concentration of the toxic indole. This resulted in improved growth and survival of the host insect. Furthermore, analysis of the genomes of gut symbiotic bacteria capable of supporting the survival of diverse stinkbug species found in nature revealed that they consistently lack the tryptophan-degrading enzyme gene. Introducing and expressing the tryptophan-degrading enzyme gene in such bacteria reduced their ability to support host growth and survival. Conversely, removing this gene from bacteria closely related to them, but possessing the tryptophan-degrading enzyme gene enhanced their symbiotic capability. These results strongly suggest that the loss of the tryptophan-degrading enzyme in bacteria may have played a crucial role in both laboratory-based experimental and natural symbiotic evolution among stinkbugs.
This study demonstrates that the loss of a single enzyme gene in gut bacteria can be a key factor in symbiotic evolution of stinkbugs. It reveals that symbiotic evolution can arise from a single mutation and elucidates important aspects of its molecular mechanism and metabolic basis. This significant achievement successfully clarifies the specific mechanisms and processes underlying the evolution of symbiosis.
Journal: Nature Microbiology
Title of paper:Tryptophanase disruption promotes insect-bacterium mutualism
Authors: Yayun Wang†, Minoru Moriyama†, Ryuichi Koga†, Kohei Oguchi, Takahiro Hosokawa, Hiroki Takai, Shuji Shigenobu, Naruo Nikoh, Takema Fukatsu (†equally contributed)
DOI: 10.1038/s41564-026-02264-z