- Similar compounds can be searched for while a query and database information remains encrypted -
Kana Shimizu (Researcher) and others, RNA Informatics Team, the Computational Biology Research Center of the National Institute of Advanced Industrial Science and Technology (AIST; President: Tamotsu Nomakuchi), in collaboration with Hiromi Arai (Postdoctoral Researcher) and others, the University of Tsukuba (President: Nobuhiro Yamada), and Kiyoshi Asai (Professor) and others, the University of Tokyo (President: Junichi Hamada), have developed a search technology for databases of compounds using secure computation.
Information on compounds used for the development of new drugs is treated as company confidential and is strictly controlled. Therefore it is difficult to send the information to external databases and conduct searches for similar compounds. Using the developed technology, a user's encrypted query and an encrypted server can be compared. Therefore, the user can search on the database while both the user and the server keep their privacy. Since the developed technology employs an efficient algorithm which is based only on an additive homomorphic cryptosystem, it enables the searches of large volumes of data at practical speed. The developed technology promotes safe and effective information exchanges between companies and is expected to contribute to open innovation in the development of new drugs.
The details of this technology will be presented at the 2011 Annual Conference of the Japanese Society for Bioinformatics / Chem-Bio Informatics Society Annual Meeting 2011 (CBI/JSBi2011 Joint Conference) to be held at the Kobe International Conference Center (Chuo-ku, Kobe) from November 8 to 10, 2011.
 |
|
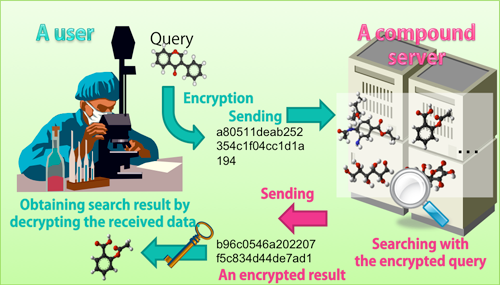
Overview of the developed system |
The field of bioinformatics requires data analysis of highly confidential data such as personal genome data and data on drug-development-related compounds. However, privacy-preserving methodology has not been discussed in depth, and the conventional remedy has been simply to isolate data from networks. Thus analyzing combinations of highly confidential information could not be conducted and researchers have been unable to make effective use of data. For example, when screening compounds having activity for drug-development targets, it is effective to search for similar compounds in databases of known compounds. However, since information on compounds is treated as company confidential and is strictly controlled, it is difficult to search on external databases. Furthermore, focused libraries offering information for a fee currently lack a facility for notifying the users beforehand whether or not an appropriate match exists for the data they hold. Naturally, many prospective users hesitate to purchase such a service for fear of wasting their investment. As a result, information vendors lose business opportunity.
Meanwhile, in the field of information science, privacy-preserving data mining has been attracting interest in recent years. In the area of privacy-preserving data mining, methods using cryptography and statistics techniques to analyze data without disclosure of confidential information are proposed.
Working from this background, the study applied the methods of privacy-preserving data mining to the field of bioinformatics, launching the development of a technology for analyzing information without disclosing highly confidential data.
In the field of bioinformatics, AIST and the University of Tokyo have been working to integrate different and separate servers, databases, and other resources. It will be increasingly necessary to handle highly confidential information, including personal genome data and data on drug-development-related compounds, in this field, so they have been investigating the importance of privacy-preserving integration of information. The University of Tsukuba has been conducting basic research into privacy-preserving data mining.
Seeking an example directly linked to an actual service, the researchers of the three parties chose as their target the problem of searching for compounds that play a vital part in the development of new drugs and devised an algorithm that employs additive homomorphic encryption. The University of Tsukuba gave advices about privacy-preserving data mining, while AIST was responsible for developing a prototype and for conducting demonstrative experiment using test data.
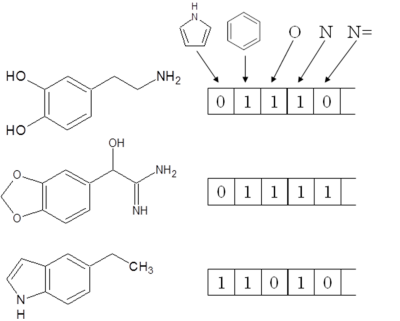
In databases etc., information of a compound is generally represented as a fixed-length bit string called a “fingerprint”. Each bit in the string shows the presence or absence of a compound characteristic. The setting “1” indicates that the characteristic is present and “0” indicates that it is absent. Figure 1 shows examples of a fingerprint. The similarity between compounds is evaluated by comparing their respective fingerprints. In general, the degree of similarity between compounds is deemed to be higher the more points at which their bit settings agree.
The most prevalent measure of similarity between fingerprints is the Tversky coefficient. In the developed technology, the Tversky coefficient was transformed into a new formula (the “decision formula”). Two fingerprints to be compared are substituted into the decision formula. If the Tversky coefficient exceeds the value set by the user, the result of the calculation is a positive value; otherwise, it is a negative value. From the calculation result of the decision formula, it is possible to determine whether or not two compounds exhibit similarity exceeding the threshold set by the user. When the compound at the user end and the compound at the server end are compared, the fingerprint for each compound undergoes additive homomorphic encryption in advance. The key used for encryption is issued by the user and transmitted to the server before the encryption is performed. Although a key for decryption is also issued by the user, it is not passed to the server end but stored. The user then transmits the encrypted fingerprint to the server end. Lacking the decryption key, the server end is unable to decrypt the content of the data sent from the user. Under these conditions, the server end computes the decision formula for the data received from the user and the data that it itself holds. As the decision formula involves addition alone, it is possible to obtain a result by computing the data encrypted using the additive homomorphic cryptosystem. Since the result obtained from the computation remains encrypted, the server end can transmit to the user the result without knowing what it is. The user decrypts the encrypted result using his/her private decryption key. By checking whether the decrypted result is a positive or negative value, the user can tell whether or not his/her query compound resembles a compound at the server end. In this way, only the user can hold the search result without disclosing compound information to the other.
 |
|
Figure 1: Examples of fingerprints |
By using these basic methods, it is possible to perform various searches according to the intended use. For example, it is possible to search a database to learn how many compounds resembling the user’s query (search data) are stored there or to return an ID-only search result for similar compounds.
Since the developed technology uses only additive homomorphic encryption, the computational cost and the volume of memory used compared with conventional methods are greatly reduced. It also allows huge amounts of data to be searched. Furthermore, the communication necessary between a user and a database server consists of only a single exchange; it is therefore possible to choose the communication means to fit the intended use. Thus, the technology is highly practical and is expected to promote safe and effective information exchange between companies and to contribute to open innovation in the drug-development field.
The already-developed software will be evaluated using real data. The results of the evaluation will serve as the basis for the improvement of the software. The researchers aim at the implementation of a compound search service in cooperation with companies.